Share this project
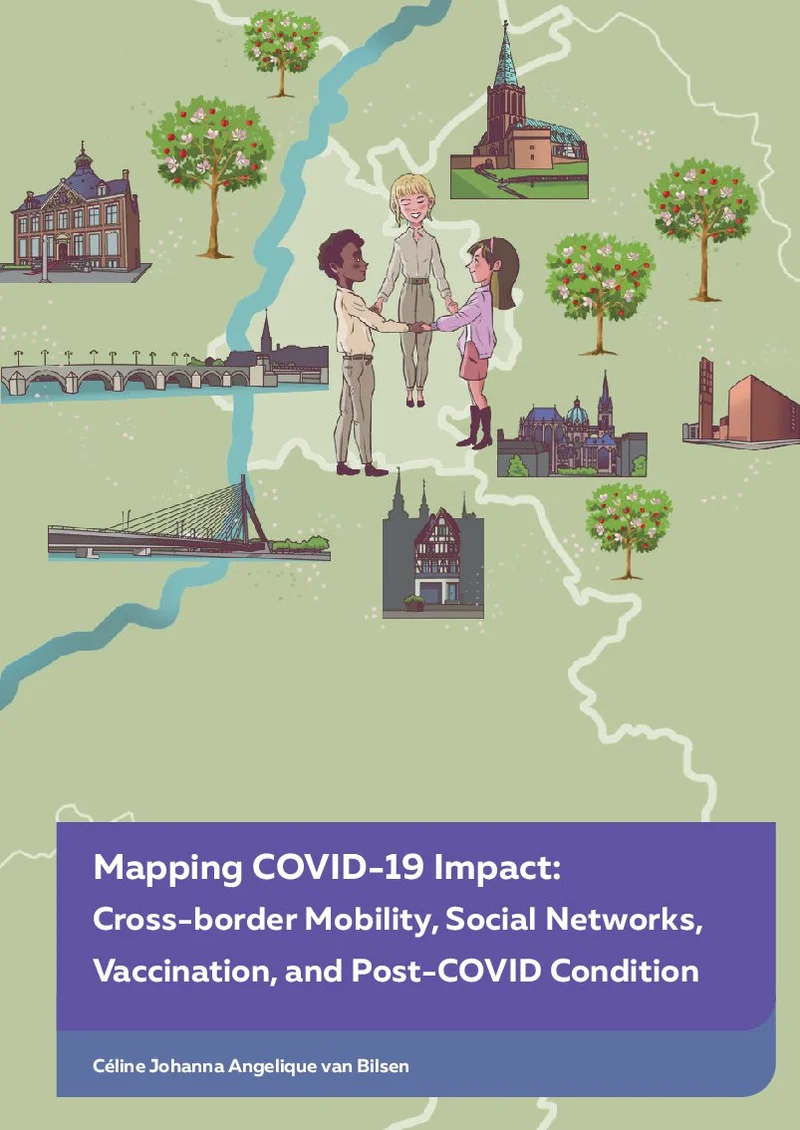
Expanding the Epi-Discovery Tool Box
Summary
The basic packaging of DNA into nucleosomes is more or less the same across the genome. However, the specific distribution of nucleosomes, their post-translational modifications, and the proteins that interact with specific chromatin structures and DNA sequences vary, leading to different types of chromatin across the genome. Each of these specific chromatin structures may have its own properties and functions. Therefore, this complexity gives rise to challenges in understanding general principles of chromatin such as the maintenance of chromatin states during transcription and DNA repair and how chromatin maintenance is coordinated spatially and temporally with DNA replication.
Chapter 1 describes how the packaging of DNA into chromatin intimately associates with the regulation of the gene expression. The packaging of chromatin into nucleosomes and the positioning of nucleosomes can help organize a gene unit such that transcription starts at the intended transcription start site. The wrapping of DNA into nucleosomes can prevent transcription initiation if the histones occupy binding sites for transcription factors or access to DNA by the basal transcription machinery. This can be modulated by post-translational modifications that can loosen histone-DNA contacts or promote histone re-assembly in the wake of RNA and DNA polymerase. During transcription and replication, histones are transiently evicted from the DNA and need to be reassembled once the process is finished. Transcription typically is associated with histone acetylation and dedicated machineries are in place to remove the acetyl groups and reset the chromatin state to prevent ectopic transcription from hyperacetylated sites.
In the presence of external signals or during cell cycle progression or development, specific gene regulatory programs need to be implemented. Such programs are initiated by the binding of regulatory proteins to specific elements at regulatory regions such enhancers and promoters and their binding is further aided by the inherently dynamic nature of chromatin which provides an opportunity for the binding of pioneering transcription factors. A cascade of binding events is then initiated, with the recruitment of chromatin remodelers that can dispel the nucleosomes and open up the chromatin. This permits the synergistic binding of general transcription factors that initiate transcription allowing the cell to implement its gene regulatory programs. While chromatin modifications regulate gene transcription, the process of transcription also changes the chromatin. Various enzymes are recruited to transcribing Pol II, for example histone acetyltransferases, ubiquitinating and de-ubiquitinating enzymes, and histone methyltransferases, which together lead to a broad spectrum of transcription-coupled histone post transcriptional modifications (PTMs). Thus, whether such modifications are a cause or consequence of transcription is an ongoing topic of interest (Henikoff & Greally, 2016; P. B. Talbert et al., 2019). Similarly, there is much interest in uncovering the underlying mechanisms that allow for the replication fork to move through chromatin and leave properly packaged daughter strands behind, without disturbing the epigenetic state of the cell. It is to be expected that chromatin modification and restoration processes are in part loci specific.
To understand the dynamics of chromatin at a detailed level, it is important to know the different players at the different loci across the genome and how chromatin states are regulated. To move towards this higher-level understanding, this thesis has focused on dissecting locus-specific chromatin regulatory pathways and protein binding events. Chapter 2 and Chapter 3, describe the application of Epi-ID to determine regulatory mechanisms, while Chapter 4 and Chapter 5 describe Epi-Decoder to decode local chromatin proteomes. These Epi-discovery techniques use chromatin immunoprecipitation followed by barcode sequencing (ChIP-Barcode-Seq) in budding yeast to probe for locus-specific chromatin regulomes and chromatin interactomes.
See also these dissertations


Structure-Preserving Data-Driven Methods for Modeling Turbulent Flows


Molecular insights into the role of VRS5 in tillering and lateral spikelet development in barley


Gamma Knife Radiosurgery for Skull Base Tumors
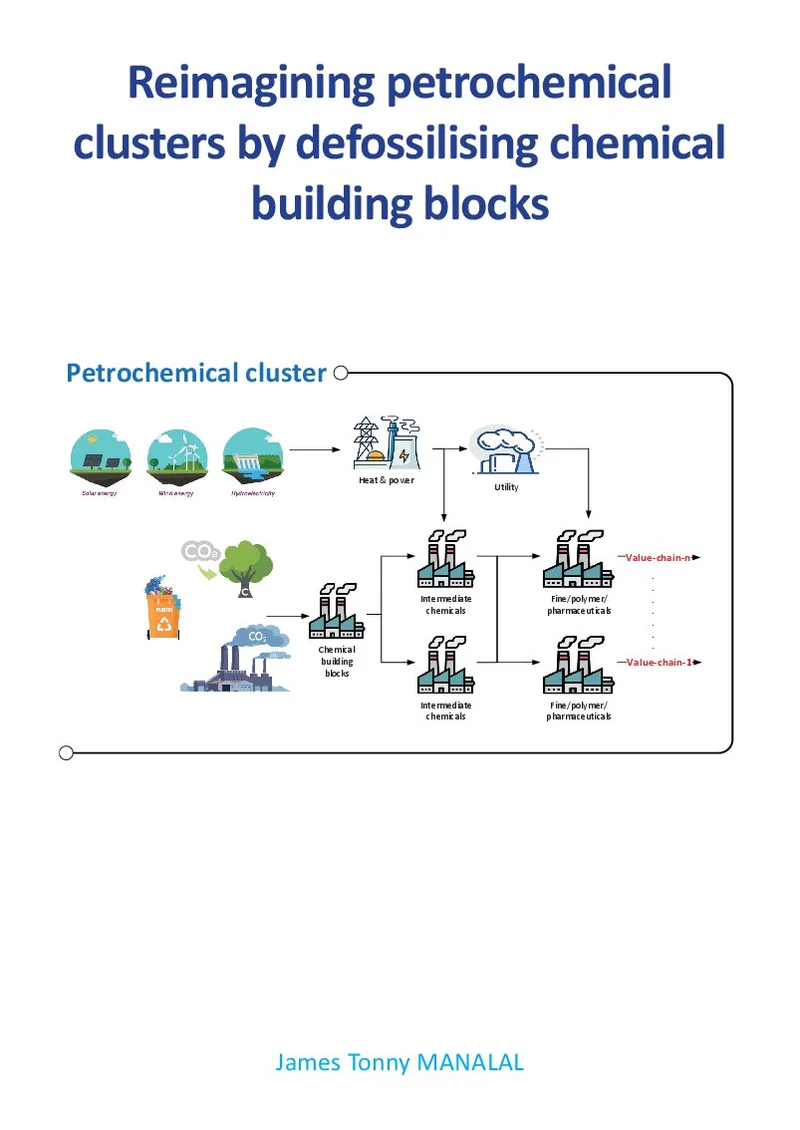

Reimagining petrochemical clusters by defossilising chemical building blocks


Microbial stabilization and protein functionality of plant-based liquids using pulsed electric fields
We print for the following universities