Share this project
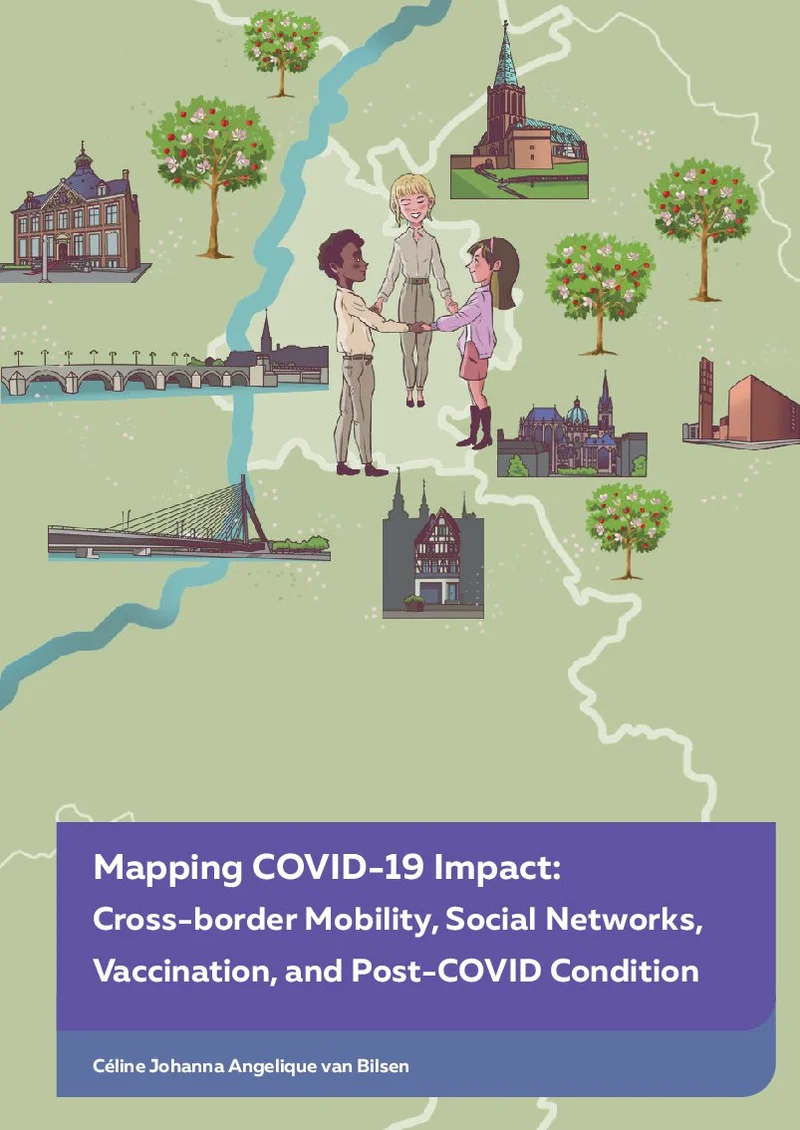
Genomic characterization and conservation of genetic diversity in cattle
Summary
Genetic diversity is the basis for livestock populations to adapt to changing environments and human demands. Conservation of genetic diversity, therefore, is an important objective of sustainable livestock production. Genetic diversity can be conserved by managing the loss of diversity within breeding programs and production systems (in situ) and by establishing gene bank collections (ex situ). Traditionally, livestock genetic diversity has been characterized and managed with pedigree-based measures of inbreeding and kinship. We now have additional opportunities to better characterize and conserve genetic diversity thanks to the increasing availability of genomic information, in particular single nucleotide polymorphism (SNP) data. These data allow, among others, to calculate more accurate coefficients of inbreeding and kinship, to study the negative effects of inbreeding on performance (‘inbreeding depression’) in more detail, and to estimate diversity across breeds. At the same time, the application of genomic information in selection schemes (‘genomic selection’) has raised questions on how to best conserve genetic diversity based on genomic information.
The overall aim of this thesis was to utilize the availability of SNP data to better characterize and conserve genetic diversity in Dutch cattle. The Holstein Friesian (HF) breed was used as the main breed of interest, because of its importance in the Dutch and global dairy cattle sector.
In Chapter 2, we used a time series of 6,280 genotyped bulls from 1986 to 2015 to investigate trends in genetic diversity in the Dutch-Flemish HF breeding program. We found major changes in diversity trends at two points in time. Around the year 2000, the introduction of optimal contribution selection (OCS) and shift in breeding goal were accompanied by a decrease in the rates of inbreeding (∆) and kinship (∆). Around 2009, the implementation of genomic selection was accompanied by an increase in ∆ and ∆. For the period 2011-2015, the ∆ and ∆ were estimated to be between 1.3% to 2.8% per generation. The observed increase in ∆ and ∆ with implementation of genomic selection was remarkable and suggests a need for stricter management of inbreeding and kinship.
In Chapters 3-5, we studied the effects of inbreeding on performance of HF cows, using pedigree, genotype and phenotype data of 38,792 first-parity cows. As expected, we observed significant inbreeding depression for yield, fertility and udder health traits. At population level, genomic inbreeding measures were found to explain more inbreeding depression than pedigree-based measures. In Chapter 3, we furthermore investigated differences in the effects of recent and ancient inbreeding. Based on pedigree, recent inbreeding appeared to be more harmful than ancient inbreeding. Especially when the pedigree-based inbreeding coefficient was split into Kalinowski’s new and ancestral components, based on whether alleles were identical-by-descent (IBD) for the first time or not, the new component was found to be more harmful than the ancestral component. Based on SNP data, we did not find such clear differences in effects of recent and ancient inbreeding. When we considered the effect of regions of homozygosity (ROH), both long ROH (which reflect recent inbreeding) and short ROH (which reflect more ancient inbreeding) contributed to inbreeding depression. While computing inbreeding coefficients we also corrected the algorithm that is commonly used to calculate Kalinowski’s coefficients (Chapter 4). In Chapter 5, we evaluated variation in inbreeding depression across the genome. We did so by estimating dominance and ROH-effects, either for one SNP at a time (through a single SNP GWAS) or for all SNPs simultaneously (through GREML with backsolving). Estimated dominance and ROH variances from GREML models were small, i.e. less than 1% of phenotypic variance. In both GWAS and GREML models, inbreeding depression appeared to be rather equally distributed across the genome and to be well captured by genome-wide homozygosity.
In Chapters 6 and 7, we demonstrated the value of the Dutch gene bank for HF and Dutch native cattle breeds. In Chapter 6, we showed that, although artificial selection has resulted in considerable genetic gains in the HF breed over time, (old) HF cryobank bulls can still be of value to today’s breeding program. First, they can be used to increase genetic diversity in the current breeding program. When we minimized relatedness with OCS, the use of genotyped cryobank bulls decreased the mean SNP-by-SNP similarity by 0.7% and the use of both genotyped and non-genotyped cryobank bulls decreased mean pedigree-based kinship by 2.6% (in absolute terms). Second, cryobank bulls can be used to increase genetic merit for a given level of diversity. We showed that the additional value of cryobank bulls was higher when relatively more emphasis was put on genetic diversity. We also found that, although the additional value of cryobank bulls was limited for the current total merit index, it was substantial for sub-indices like fertility. Anticipating changes in breeding goals in future, we concluded that the gene bank collection is a valuable resource. In Chapter 7, we characterized genetic diversity in the gene bank across Dutch native cattle breeds. Besides the HF breed, 7 Dutch native cattle breeds are stored in the gene bank. We showed that, based on SNP data, Dutch native breeds genetically differed from the HF breed, suggesting they harbor unique genetic diversity. Among Dutch native breeds, there was admixture. Consequently, when we set up a ‘core set’ in which the expected heterozygosity was maximized through OCS, some breeds were assigned low contributions. Overall, our results show that gene bank collections are valuable resources, not only for small local breeds, but also for large commercial breeds.
In Chapter 8, the general discussion, I further addressed some major questions and opportunities for genomic management of genetic diversity. I showed that the increase in ∆ and ∆ with genomic selection in HF is a global trend and argued that the increase is not necessarily due to the methodology, but rather to a change in system. In addition, I discussed why the SNP-by-SNP similarity is an important measure for conservation of genetic diversity. Moreover, I discussed how OCS can be extended, providing additional conservation opportunities (e.g. for region-specific diversity management), and described the benefits of using sequence data. I also discussed how characterization of gene bank collections may enhance their utilization and how gene bank collections are expected to move towards bio-digital resource centers. Finally, I emphasized the importance of conservation of genetic diversity by discussing expected changes in future breeding goals.
See also these dissertations


Structure-Preserving Data-Driven Methods for Modeling Turbulent Flows


Molecular insights into the role of VRS5 in tillering and lateral spikelet development in barley


Gamma Knife Radiosurgery for Skull Base Tumors

Reimagining petrochemical clusters by defossilising chemical building blocks


Microbial stabilization and protein functionality of plant-based liquids using pulsed electric fields
We print for the following universities